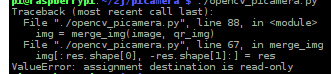
Alibaba cloud CentOS FTP server configures FTP in passive mode and reports an error
200 port command successful. Consider using PASV. 425 failed to establish connection
Answer:
you are using FTP in active mode.
Due to firewall and NAT, it may be troublesome to set up FTP in active mode nowadays.
The server may not be able to connect back to the client to establish a data transfer connection, which may be due to your local firewall or NAT.
Or your client does not know its external IP address, but provides an internal address to the server (in the
port
command), and the server obviously cannot use the internal address. But this is not the case, because vsftpd refuses the source address (
port) of FTP control connection by default_ Promiscuous
instruction).
Please refer to my article “active mode network configuration”.
If possible, you should use passive mode, as it usually does not require other settings on the client side. This is what the server suggests to you through “consider PASV.”. This
PASV
is the FTP command used to enter passive mode.
Unfortunately, the windows FTP command line client (
0 ftp.exe
)Passive mode is not supported at all. Now, it’s useless.
Please use any other third party windows FTP command line client instead. Most others support passive mode.
For example, the winscp FTP client defaults to passive mode, and provides guidelines for converting windows FTP scripts to winscp scripts.
(I’m the author of winscp)
Answer:
in fact, your window firewall blocks the connection, so you need to enter these commands from the administrator cmd.exe .
1) Netsh advfirewall add rule name = “FTP” dir = executing = allow program =% systemroot% \ \ system32\ ftp.exe Enable = yes protocol = TCP
2) Netsh advfirewall add rule name = “FTP” dir = executing = allow program =% systemroot% \ \ system32\ ftp.exe Enable = yes protocol = UDP
If something goes wrong, you can recover in the following ways:
1) Netsh advfirewall delete rule name = “FTP” program =% systemroot% \ \ system32\ ftp.exe
Questions:
I have setup FTP server in Ubuntu 12.04 LTS.
Now when when I try to connect to FTP server from Windows 7 through command-line
ftp.exe
, I get successfully connected but I cannot get the list of directory. I get error
200 PORT command successful. Consider using PASV.
425 Failed to establish connection.
Answers:
Try using the
passive
command before using
ls
.
From FTP client, to check if the FTP server supports passive mode, after login, type
quote PASV
.
Following are connection examples to a vsftpd server with passive mode on and off
vsftpd
with
pasv_ enable=NO
:
ftp localhost
Connected to localhost.localdomain .
220 (vsFTPd 2.3.5)
Name ( localhost:john ): anonymous
331 Please specify the password.
Password:
230 Login successful.
Remote system type is UNIX.
Using binary mode to transfer files.
ftp> quote PASV
550 Permission denied.
ftp>
vsftpd
with
pasv_ enable=YES
:
ftp localhost
Connected to localhost.localdomain .
220 (vsFTPd 2.3.5)
Name ( localhost:john ): anonymous
331 Please specify the password.
Password:
230 Login successful.
Remote system type is UNIX.
Using binary mode to transfer files.
ftp> quote PASV
227 Entering Passive Mode (127,0,0,1,173,104).
ftp>
Answers:
You are using the FTP in an active mode.
Setting up the FTP in the active mode can be cumbersome nowadays due to firewalls and NATs.
It’s likely because of your local firewall or NAT that the server was not able to connect back to your client to establish data transfer connection.
Or your client is not aware of its external IP address and provides an internal address instead to the server (in
PORT
command), which the server is obviously not able to use. But it should not be the case, as vsftpd by default rejects data transfer address not identical to source address of FTP control connection (the
port_ promiscuous
directive).
See my article Network Configuration for Active Mode.
If possible, you should use a passive mode as it typically requires no additional setup on a client-side. That’s also what the server suggested you by “Consider using PASV”. The
PASV
is an FTP command used to enter the passive mode.
Unfortunately Windows FTP command-line client (the
ftp.exe
) does not support passive mode at all. It makes it pretty useless nowadays.
Use any other 3rd party Windows FTP command-line client instead. Most other support the passive mode.
For example WinSCP FTP client defaults to the passive mode and there’s a guide available for converting Windows FTP script to WinSCP script.
(I’m the author of WinSCP)
Answers:
Actually your window firewall blocking the connection so you need to Enter these commands into cmd.exe from Administrator.
netsh advfirewall firewall add rule name=”FTP” dir=in action=allow program=%SystemRoot%\System32\ ftp.exe enable=yes protocol=tcp netsh advfirewall firewall add rule name=”FTP” dir=in action=allow program=%SystemRoot%\System32\ ftp.exe enable=yes protocol=udp
if in case something goes wrong then you can revert by this:
netsh advfirewall firewall delete rule name=”FTP” program=%SystemRoot%\System32\ ftp.exe